Optional: Step 7: Identify the single best trendline
Remember how you chose several trendlines in “Process Data”, resulting in all of these (or similar) output folders?


If “Process Data” is completed and you have these folders, you can use the “Best Trendline ID” tool to identify the single best performing trendline. This may be helpful if you can’t spend lots of time on processing data, and you want to optimize which single trendline performs best for your protein.
7.1 Open Best Trendline ID
Open the tool over the main interface
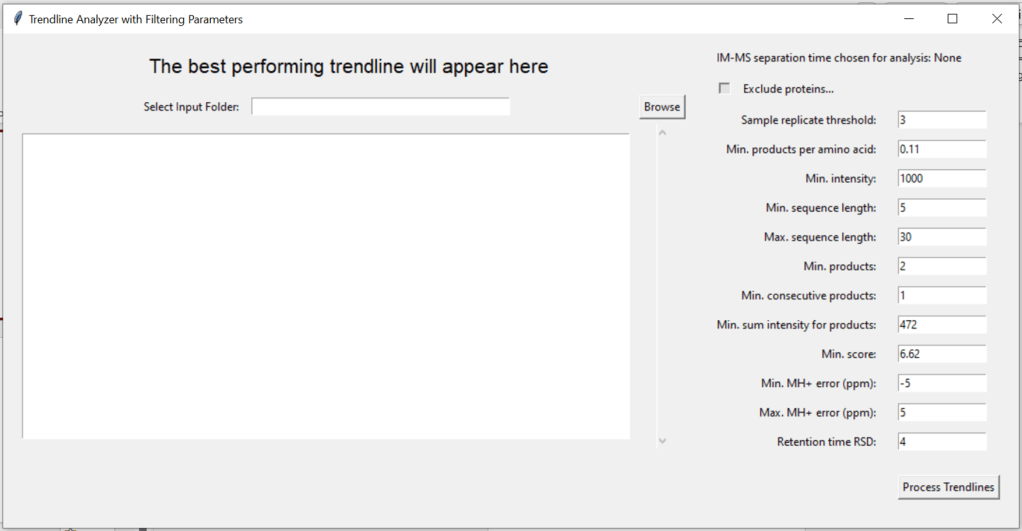
7.2. Select an input folder
Browse for the folder that contains the processed data from “Process Data”. It may look like this:

7.3 Select a IM-MS separation time
After selecting the input folder, a window will pop up that shows all IM-MS separation times that were found. Select one.

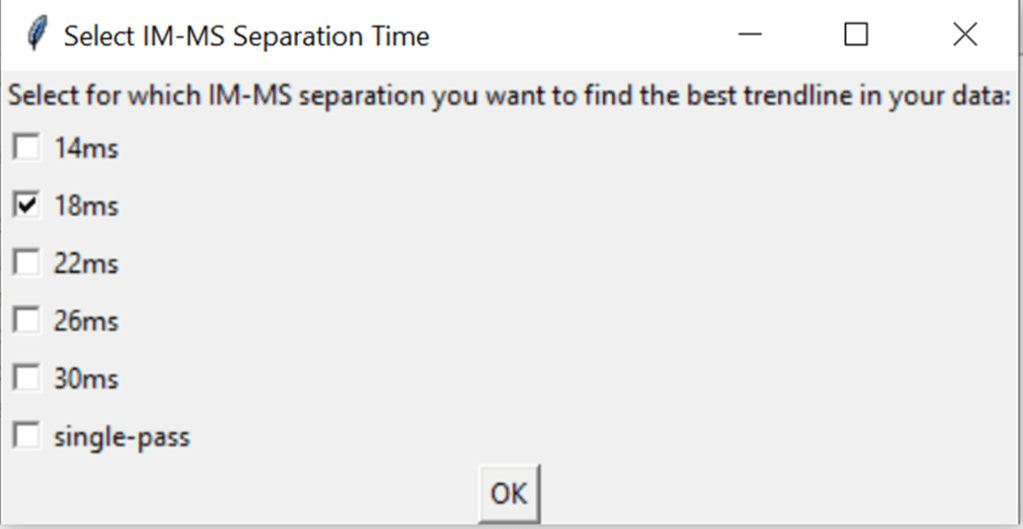

Hint: it uses the file names to know which IM-MS separation time was used, e.g. 120923_11_09_2024_PhosB_18ms_IA_final_peptide will get counted as 18ms. If a file is in the sub-folder 1_1 and has no separation time, the software assumes that this is the single-pass data, and lists it as such.
7.4 Choose which (if any) proteins to exclude
Select proteins you want to exclude from processing by unticking them
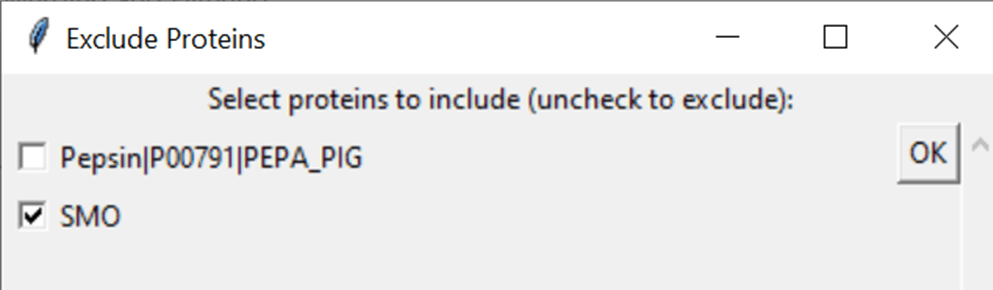

7.5 Fill out the filtering parameters
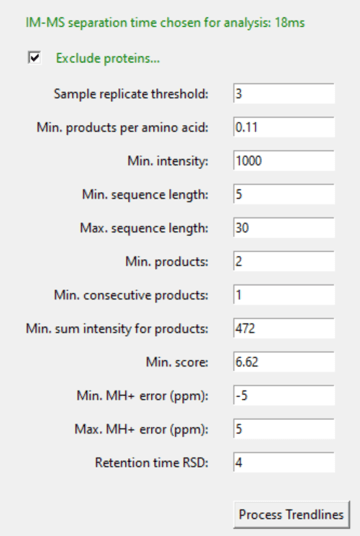
7.6 Click “Process Trendlines” & look at the results
Processing the files should take less than a minute. You can look at the results by scrolling to the bottom of the user interface; the results of the single highest performing trendline will also get saved with a timestamp in a “processed results” folder:
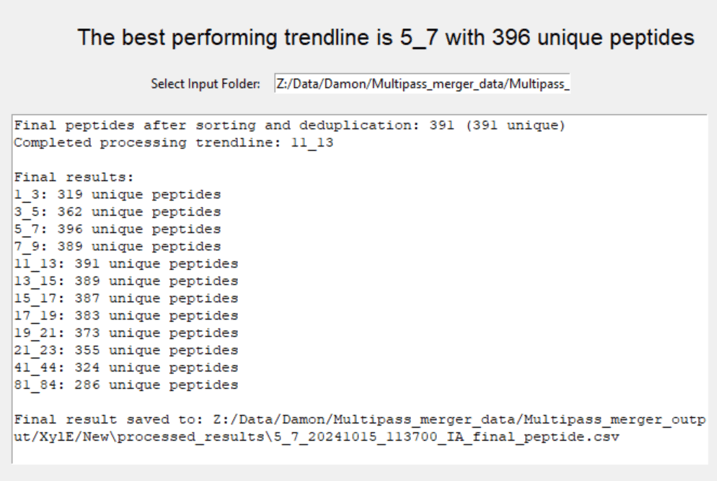
Next time you process data for this protein, you will know that e.g. trendline 5-7 performs best (=smoothens the data best) for the 18ms multi-pass separation. Although your results will not be as good as if you would have merged several trendline results together, using only 1 trendline will save you a lot of processing time.
